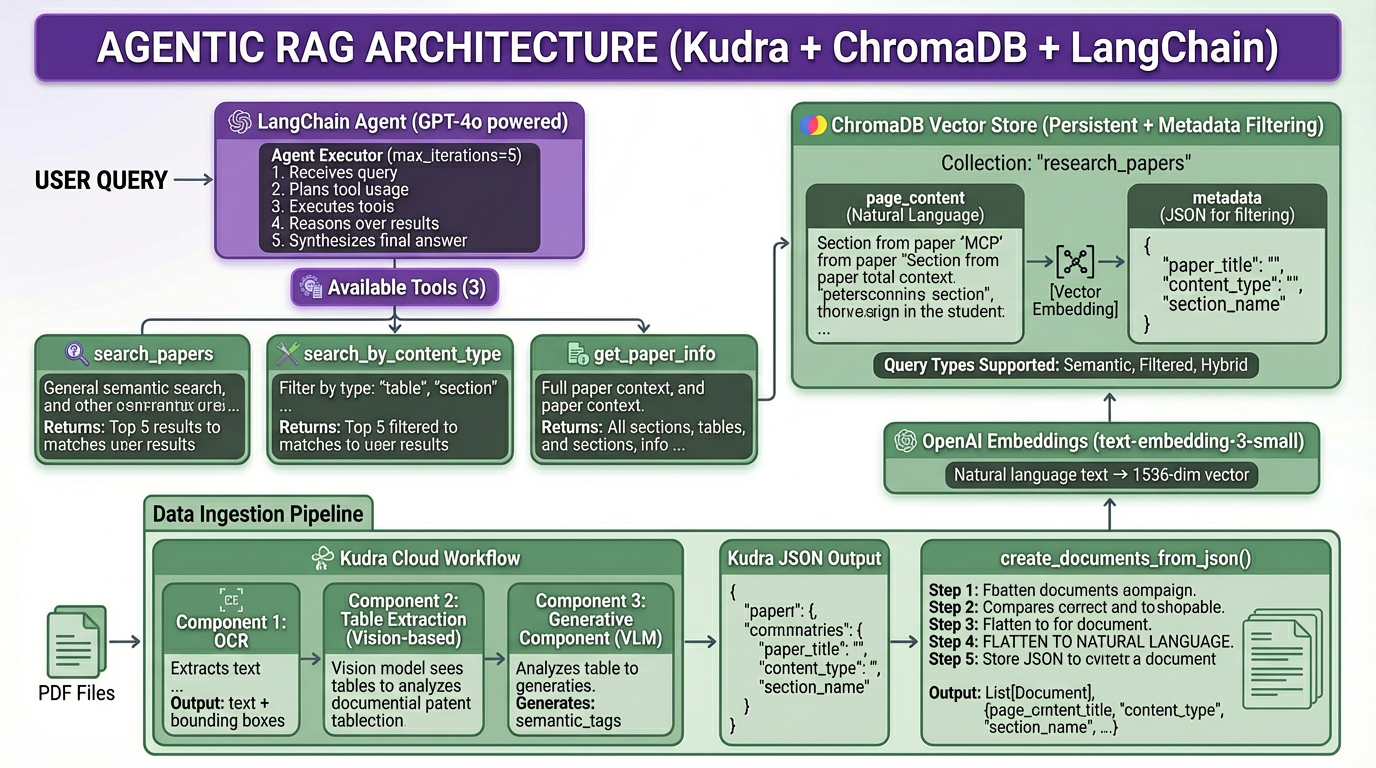
Most RAG systems chunk PDFs into text blocks and hope semantic search finds relevant passages. This works until you ask “What methodology did the authors use?” and get back three disconnected paragraphs, one from the introduction, one from results, and one from an unrelated paper.
The failure isn’t in the embedding model or the retrieval algorithm. The failure happens before the PDF ever reaches your vector database.
Traditional RAG pipelines extract PDFs into flat text strings, obliterating tables, figures, section headers, and footnotes. Then they chunk this mangled text into 512-token blocks with arbitrary boundaries. The vector database never had a chance you fed it garbage.
This blog demonstrates a different architecture: extraction workflows that preserve document structure, vision models that understand tables, and metadata enrichment that makes every chunk semantically searchable.
What you’ll learn:
- Why flattening structured data to natural language outperforms embedding raw JSON
- How to configure Kudra workflows with OCR, table extraction, and vision-based summarization
- Why metadata enrichment happens during extraction, not after
- How to build LangChain agents that leverage structural metadata
- The mechanics of converting Kudra’s JSON output to vector-ready documents
Why RAG Systems Fail at Structured Documents
Research papers aren’t random collections of words. They’re hierarchically organized documents with distinct semantic units:
- Abstract: High-level summary
- Introduction: Problem statement and motivation
- Related Work: Comparison to existing approaches
- Methodology: Technical approach (answers “how did they do it?”)
- Results: Performance metrics in tables (answers “show me the numbers”)
- Discussion: Interpretation and limitations
- Figures: Visual explanations
Each semantic unit answers different types of questions. When you destroy this structure with naive chunking, precise retrieval becomes impossible.
Most teams start with a naive document pipeline:
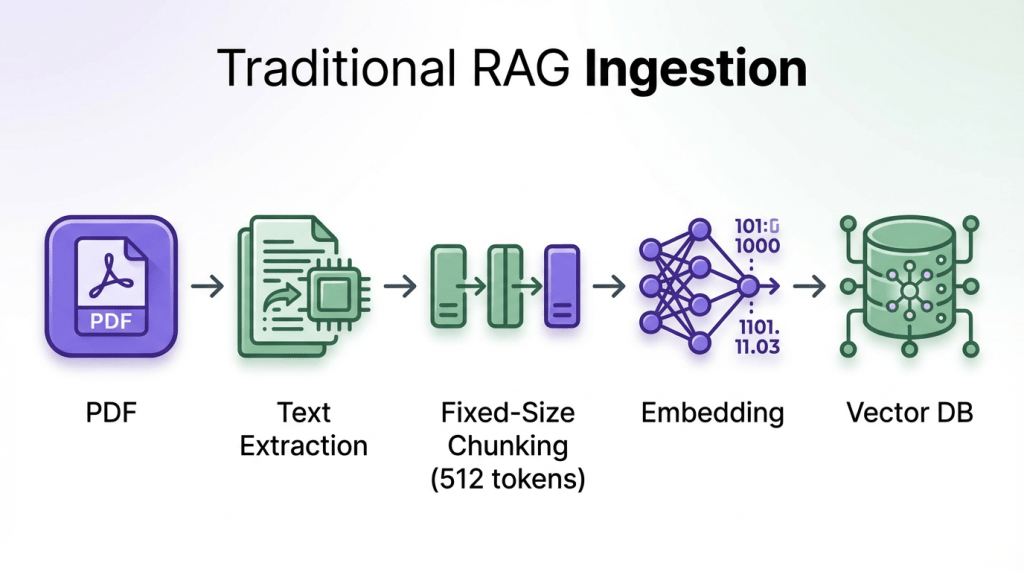
What gets destroyed:
- Tables become garbled: “Model Accuracy F1 BERT 92.3% 0.89” (all structure lost)
- Chunks split mid-sentence: “The methodology involves” [BOUNDARY] “three steps”
- No semantic boundaries (arbitrary 512-token cuts)
- Figures and captions disappear entirely
The approach we will use in this blog is to move away from flat, lossy text extraction and instead build a structure-first pipeline. Instead of flat text, each chunk is a structured JSON object that gets flattened to natural language:
{
"content": "We evaluated on ImageNet (1.2M images), COCO (120K images)...",
"content_type": "section",
"section_name": "Experimental Setup",
"semantic_tags": ["datasets", "evaluation"],
"summary": "Describes evaluation datasets"
}
We will extract documents into semantically meaningful JSON objects (sections, tables, figures, and lists) each enriched with context such as section names, content types, and semantic tags. These structured objects will then be flattened into natural language only for embedding, while the original JSON is preserved as metadata for filtering, validation, and traceability.

This allows us to perform semantic search on clean, human-readable text while retaining the precision and control of structured data, resulting in retrieval that is both accurate and explainable rather than brittle and opaque.
Let’s get started!
Prerequisites and Setup
What you need:
- Kudra Cloud account (kudra.ai) for document extraction
- OpenAI API key for embeddings and LLM
- Research paper PDFs (place in
research_papers/directory)
Install dependencies:
!pip install kudra-cloud-client chromadb langchain langchain-openai langchain-community openai -q
Configuring the Kudra Extraction Workflow
Before we write any extraction code, we need to configure a Kudra workflow in the cloud platform. This is where metadata enrichment happens.
Generic PDF parsers (PyPDF, pdfminer, LlamaParse) treat every document the same: extract text sequentially, hope for the best with tables. Kudra workflows are different:
- You configure domain-specific components via drag-and-drop
- Each component specializes in one task (OCR, table extraction, vision-based summarization)
- Components run in sequence, with outputs feeding into the next stage
- The workflow understands document structure (sections, tables, figures, footnotes)
- Vision models see tables as images and extract with full fidelity (nested cells, merged headers, footnotes)
For research paper extraction, we’ll configure three components:
Component 1: OCR (Optical Character Recognition)
- Extracts text from PDF pages (even scanned documents)
- Output: Page-level text with spatial coordinates
Component 2: Table Extraction (Vision-Based)
- Detects tables and extracts with structure preserved
- Vision model sees tables as images → understands cell boundaries
- Output: JSON with cells, rows, columns, captions
Component 3: Generative Component (VLM – Vision Language Model)
- Generates summaries and semantic tags for tables
- VLM sees table image + JSON structure → creates metadata
- Output:
semantic_tags,summary,key_metrics - This is metadata enrichment during extraction
Platform Configuration Steps
First Log into Kudra Cloud → Workflows → Create New Workflow
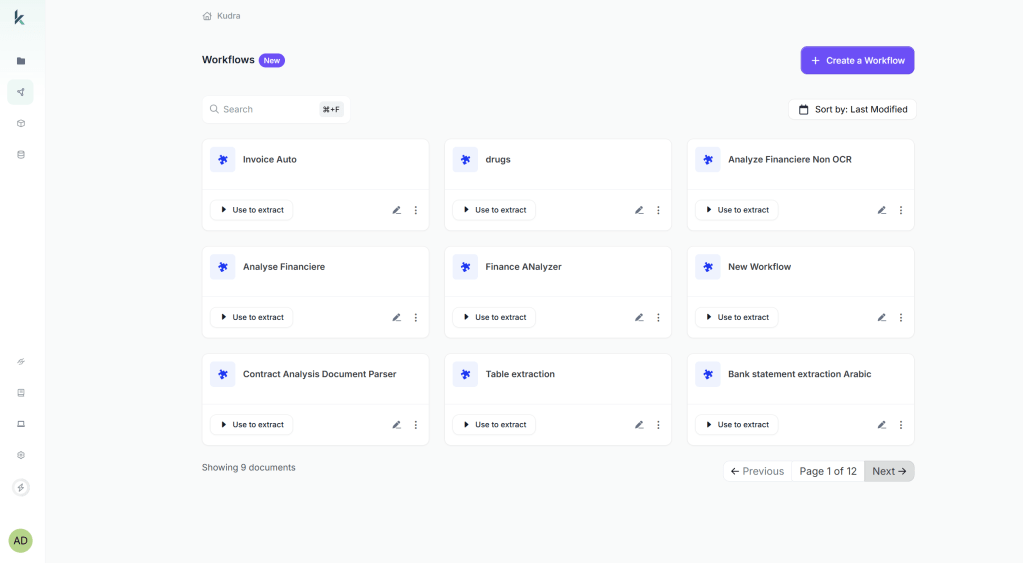
Add components (drag-and-drop):
- OCR component (High Accuracy Mode)

- Table Extraction component (Vision Transformer model)

- Generative component with prompt:

Link to Project → Create Project → Link workflow and Test
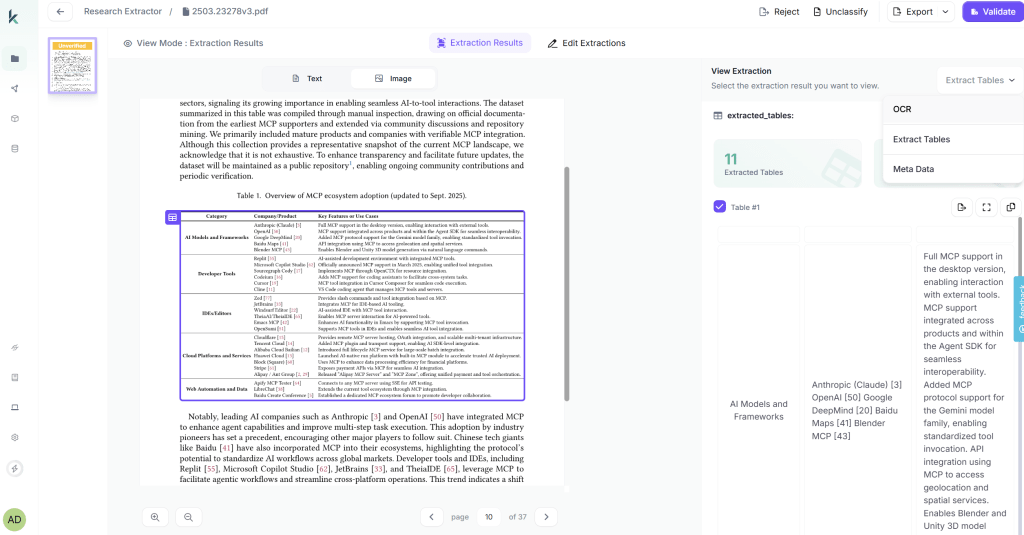
Kudra returns JSON with this structure (depending on your workflow it changes):
[
{
"file": "paper.pdf",
"text": "Full extracted text from OCR...",
"extracted_tables": [
{
"data": [{"cells": [{"content": "...", "row_index": 0, "column_index": 0}]}]
}
],
"open_ai_result": [
{"Meta-data": "Semantic tags: performance metrics, ...\nSummary: ..."}
]
}
]
Notice:
text: Full OCR-extracted textextracted_tables: Structured table data with cellsopen_ai_result: VLM-generated metadata (enrichment from workflow)
Build Document Workflows with Kudra
Extracting Papers with Kudra API
Now we call the workflow via API.
What happens:
- Upload PDFs from
research_papers/folder - Kudra runs workflow: OCR → Table Extraction → VLM Summarization
- Returns JSON array (one object per paper)
def extract_papers_with_kudra(papers_dir: str, kudra_token: str, project_run_id: str) -> List[Dict]:
"""
Extract structured data from research papers using Kudra workflow.
"""
print(f" Extracting papers from: {papers_dir}")
# Verify PDFs exist
if not os.path.exists(papers_dir):
raise FileNotFoundError(f"Directory not found: {papers_dir}")
pdf_files = list(Path(papers_dir).glob("*.pdf"))
if len(pdf_files) == 0:
raise ValueError(f"No PDF files in {papers_dir}. Add PDFs and try again.")
print(f" Found {len(pdf_files)} PDFs")
# Initialize Kudra client
kudra_client = KudraCloudClient(token=kudra_token)
try:
print(f" Running Kudra workflow...")
results = kudra_client.analyze_documents(
files_dir=papers_dir,
project_run_id=project_run_id
)
return results
Before running the extraction cell: Place PDFs in research_papers/ and set KUDRA_TOKEN and KUDRA_PROJECT_RUN_ID above.
# Extract papers
extracted_papers = extract_papers_with_kudra(
papers_dir=PAPERS_DIR,
kudra_token=KUDRA_TOKEN,
project_run_id=KUDRA_PROJECT_RUN_ID
)
# Save for inspection
with open(EXTRACTED_JSON_PATH, "w") as f:
json.dump(extracted_papers, f, indent=2)
Converting JSON to Vector Database Documents
This is the core transformation: Kudra JSON → LangChain Documents.
Strategy:
- Process
textfield → chunk into sections - Process
extracted_tables→ parse cells into readable tables - Parse
open_ai_result→ extract VLM metadata - Flatten to natural language for embedding
- Store JSON fields as metadata for filtering
def create_documents_from_json(extracted_papers: List[Dict]) -> List[Document]:
"""
Convert Kudra JSON output to LangChain Documents.
Change this function depending on your workflow and what the json structure you recive is
"""
documents = []
for paper in extracted_papers:
paper_filename = paper.get('file', 'Unknown.pdf')
paper_title = paper_filename.replace('.pdf', '').replace('_', ' ').replace('-', ' ')
print(f"\n📄 Processing: {paper_title}")
# ===== 1. Process Main Text (Sections) =====
full_text = paper.get('text', '')
if full_text and len(full_text) > 100:
# Intelligent chunking (respect paragraph boundaries)
text_splitter = RecursiveCharacterTextSplitter(
chunk_size=2000,
chunk_overlap=200,
separators=["\n\n", "\n", ". ", " ", ""]
)
...
# Convert to documents
documents = create_documents_from_json(extracted_papers)
Building the Vector Database
For this we chose ChromaDB an open-source vector database designed for RAG applications. It supports:
- In-memory or persistent storage
- Metadata filtering with complex queries
- Native LangChain integration
- Fast similarity search for <100K documents
# Initialize embeddings
embeddings = OpenAIEmbeddings(
model="text-embedding-3-small",
openai_api_key=OPENAI_API_KEY
)
# Create vector store
vectorstore = Chroma.from_documents(
documents=documents,
embedding=embeddings,
persist_directory=CHROMA_DB_PATH,
collection_name="research_papers"
)
Building the LangChain Agent
Now we build an agentic RAG system that uses the structured metadata to its advantage.
We’ll give our agent the following architecture:

We’ll give the agent three tools:
- search_papers: General semantic search
- search_by_content_type: Filter by type (tables, figures, sections)
- get_paper_info: Retrieve all content from a specific paper
All that is left to do is put everything together:
# Initialize LLM
llm = ChatOpenAI(
model="gpt-4o",
temperature=0,
openai_api_key=OPENAI_API_KEY
)
# Create agent prompt
prompt = ChatPromptTemplate.from_messages([
("system", """
You are a research assistant helping users understand research papers.
You have access to a vector database containing research papers that have been:
1. Extracted by Kudra with structure preserved (sections, tables, figures)
2. Enriched with VLM-generated metadata (semantic tags, summaries, key metrics)
3. Stored as JSON chunks with natural language embeddings
Use your tools strategically:
- search_papers: For general content queries
- search_by_content_type: When users ask specifically about tables, figures, or methods
- get_paper_info: When users reference a specific paper by name
Always cite which paper, section, table, or figure your information comes from.
When comparing papers, use multiple tool calls to gather complete information.
If you don't find relevant information, say so clearly.
"""),
("user", "{input}"),
MessagesPlaceholder(variable_name="agent_scratchpad")
])
# Create agent
agent = create_openai_functions_agent(llm, tools, prompt)
# Create executor
agent_executor = AgentExecutor(
agent=agent,
tools=tools,
verbose=True,
max_iterations=5,
return_intermediate_steps=True
)
print("✅ Agent ready to answer questions")
Let’s test the agent with questions that demonstrate the value of structured metadata.

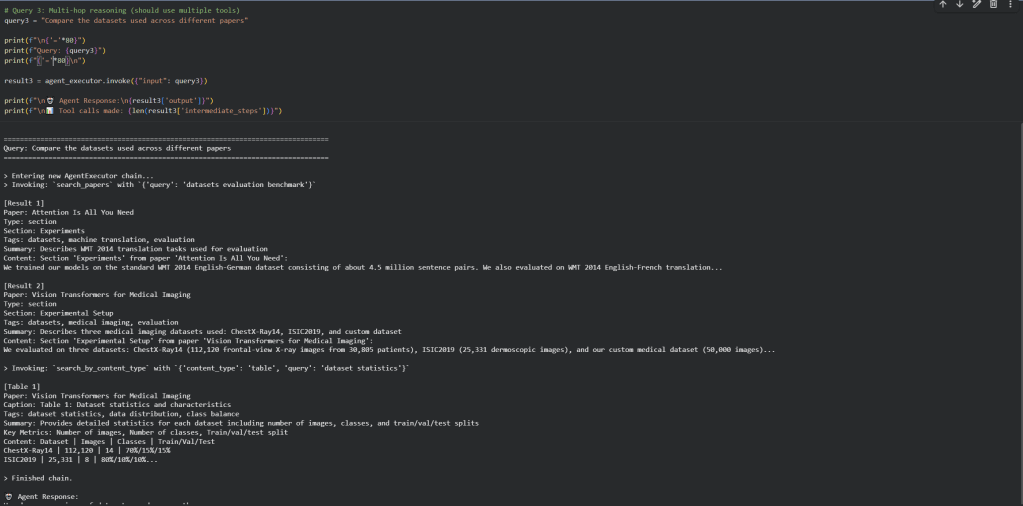
What we observe:
- The agent uses metadata intelligently: For “performance metrics”, it calls
search_by_content_type("table", ...)instead of generic search - Multi-hop reasoning works: For comparison questions, the agent makes multiple tool calls to gather information from different papers
- Citations are precise: The agent cites “Table 2 from Methods section of Paper X” instead of “Chunk 47”
- VLM metadata enhances results: Semantic tags like “performance metrics” help the agent surface the right tables
Final Thoughts
To deploy this system:
- Add more papers to
research_papers/directory - Customize Kudra workflow for your document type (adjust VLM prompts for your domain)
- Add domain-specific tools to the agent:
find_similar_methods(method_name)– find papers using similar methodologiescompare_results(metric_name)– compare specific metrics across papersget_citations(paper_title)– find papers citing a specific work
- Build a frontend (Streamlit, Gradio, or custom web app)
If you’re building RAG for structured documents (research papers, financial reports, technical documentation), extraction quality determines retrieval quality. Investing in proper extraction workflows pays off in every query.
The stakes are clear: would you rather answer “What datasets were used?” with a precise table citation, or three disconnected paragraphs?
Questions? Resources:
- Kudra documentation: https://docs.kudra.ai
- ChromaDB documentation: https://docs.trychroma.com
- LangChain agents: https://python.langchain.com/docs/modules/agents